 (1962) Host specificity of DNA produced by Escherichia coli. (1953) Host controlled variation in bacterial viruses. (1952) A nonhereditary, host-induced variation of bacteria viruses. (1995) Restriction and modification systems of Neisseria gonorrhoeae. and Roberts, R.J., eds., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY (1993) In: Nucleases, Second Edition Linn, S.M., Lloyd, S.R. Because of the ability of these enzymes to cleave DNA at specific recognition sites, they have continued to play a fundamental role in cloning and DNA typing applications. Smith and Daniel Nathans shared the 1978 Nobel Prize for Medicine and Physiology for their discovery of restriction enzymes and their application to molecular genetics. In 1970, Smith and colleagues described the purification of the first Type II restriction enzyme, Hind II (9), and the characterization of its recognition and cleavage site (10). These were later classified as Type I restriction enzymes, which cleave DNA at random positions, often far removed from the recognition site. In 1968, Arber and Linn demonstrated nuclease activity of Eco B restriction enzyme (7) and Meselson and Yuan purified a similar enzyme from E. This modified DNA is able to successfully replicate and infect the second host, but since that host does not contain the same modification system as the first, the modified phage lose their ability to replicate on the original host. However, a small proportion of the phage DNA is modified prior to degradation by the endonuclease. Unmodified (foreign) DNA, such as that of an infecting phage, is degraded by the endonuclease, restricting phage infection (hence the term restriction endonuclease). They postulated that certain bacterial strains contain an endonuclease that is able to cleave DNA, and that some strains contain a strain-specific modification system that is responsible for protecting host DNA from the action of its own endonuclease. Phage isolated from these plaques were able to re-infect the second strain and grow well, but lost the ability to grow on the original strain.Īrber and Dussoix (5, 6)proposed a molecular model to explain host-controlled restriction modification. 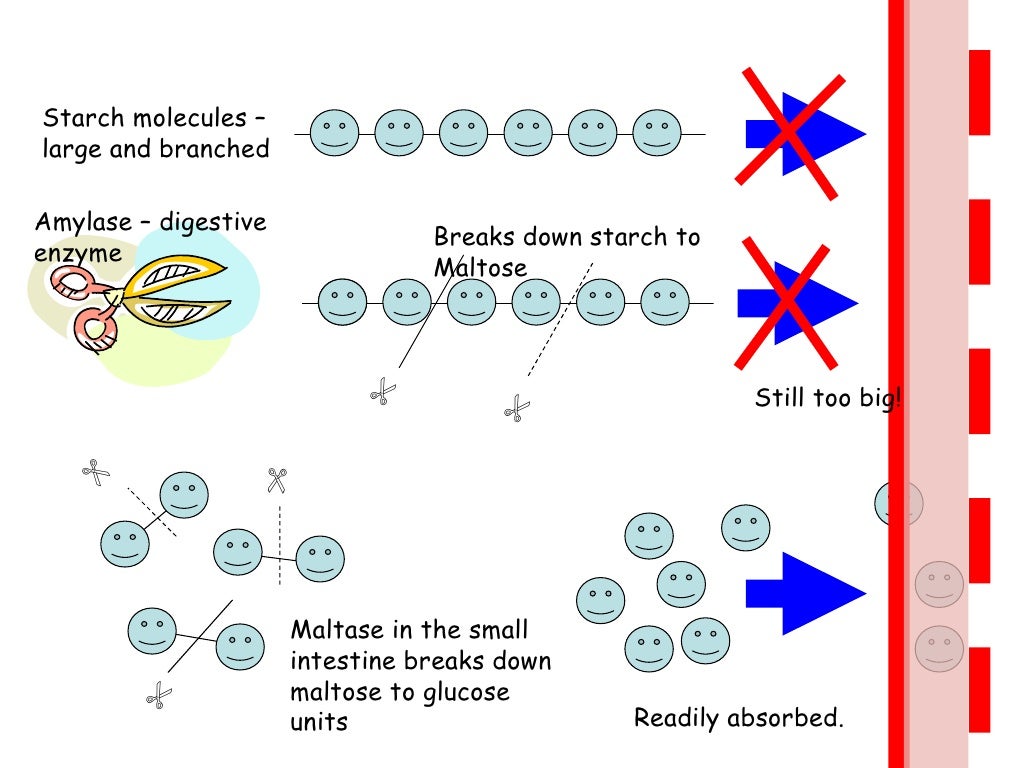
They observed that bacteriophage that grew well in one bacterial strain often grew poorly in a second, forming only a few plaques. In the early 1950s, Luria and colleagues (3, 4) reported a phenomenon known as host-controlled restriction modification. It has been estimated that 25% of all bacteria contain at least one restriction enzyme (1) and as many as 7 have been found in a single species (2) They have been found mostly in bacteria, but have also been isolated from viruses, archaea and eukaryotes. Approximately 3,000 restriction enzymes, recognizing over 230 different DNA sequences, have been discovered.  
Restriction enzymes recognize short DNA sequences and cleave double-stranded DNA at specific sites within or adjacent to these sequences. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
